3 Attribute data operations
Prerequisites
- This chapter requires the following packages to be installed and attached:
library(sf) # vector data package introduced in Chapter 2
library(terra) # raster data package introduced in Chapter 2
library(dplyr) # tidyverse package for data frame manipulation- It relies on spData, which loads datasets used in the code examples of this chapter:
- Also ensure you have installed the tidyr package, or the tidyverse of which it is a part, if you want to run data ‘tidying’ operations in Section 3.2.5.
3.1 Introduction
Attribute data is non-spatial information associated with geographic (geometry) data.
A bus stop provides a simple example: its position would typically be represented by latitude and longitude coordinates (geometry data), in addition to its name.
The Elephant & Castle / New Kent Road stop in London, for example has coordinates of \(-0.098\) degrees longitude and 51.495 degrees latitude which can be represented as POINT (-0.098 51.495) in the sfc representation described in Chapter 2.
Attributes, such as name, of the POINT feature (to use simple features terminology) are the topic of this chapter.
Another example is the elevation value (attribute) for a specific grid cell in raster data. Unlike the vector data model, the raster data model stores the coordinate of the grid cell indirectly, meaning the distinction between attribute and spatial information is less clear. To illustrate the point, think of a pixel in row 3 and column 4 of a raster matrix. Its spatial location is defined by its index in the matrix: move from the origin four cells in the x direction (typically east and right on maps) and three cells in the y direction (typically south and down). The raster’s resolution defines the distance for each x- and y-step which is specified in a header. The header is a vital component of raster datasets which specifies how pixels relate to spatial coordinates (see also Chapter 4).
This chapter teaches how to manipulate geographic objects based on attributes such as the names of bus stops in a vector dataset and elevations of pixels in a raster dataset.
For vector data, this means techniques such as subsetting and aggregation (see Sections 3.2.1 to 3.2.3).
Sections 3.2.4 and 3.2.5 demonstrate how to join data onto simple feature objects using a shared ID and how to create new variables, respectively.
Each of these operations has a spatial equivalent:
the [ operator in base R, for example, works equally for subsetting objects based on their attribute and spatial objects; you can also join attributes in two geographic datasets using spatial joins.
This is good news: skills developed in this chapter are cross-transferable.
After a deep dive into various types of vector attribute operations in the next section, raster attribute data operations are covered. Creation of raster layers containing continuous and categorical attributes and extraction of cell values from one or more layer (raster subsetting) (Section 3.3.1) are demonstrated. Section 3.3.2 provides an overview of ‘global’ raster operations which can be used to summarize entire raster datasets. Chapter 4 extends the methods presented here to the spatial world.
3.2 Vector attribute manipulation
Geographic vector datasets are well supported in R thanks to the sf class, which extends base R’s data.frame.
Like data frames, sf objects have one column per attribute variable (such as ‘name’) and one row per observation or feature (e.g., per bus station).
sf objects differ from basic data frames because they have a geometry column of class sfc which can contain a range of geographic entities (single and ‘multi’ point, line, and polygon features) per row.
This was described in Chapter 2, which demonstrated how generic methods such as plot() and summary() work with sf objects.
sf also provides generics that allow sf objects to behave like regular data frames, as shown by printing the class’s methods:
methods(class = "sf") # methods for sf objects, first 12 shown
#> [1] [ [[<- $<- aggregate
#> [5] as.data.frame cbind coerce filter
#> [9] identify initialize merge plot Many of these (aggregate(), cbind(), merge(), rbind() and [) are for manipulating data frames.
rbind(), for example, binds rows of data frames together, one ‘on top’ of the other.
$<- creates new columns.
A key feature of sf objects is that they store spatial and non-spatial data in the same way, as columns in a data.frame.
The geometry column of sf objects is typically called geometry or geom, but any name can be used.
The following command, for example, creates a geometry column named g:
st_sf(data.frame(n = world$name_long), g = world$geom)
wkb_geometry and the_geom.
sf objects can also extend the tidyverse classes for data frames, tbl_df and tbl.
Thus sf enables the full power of R’s data analysis capabilities to be unleashed on geographic data, whether you use base R or tidyverse functions for data analysis.
sf objects can also be used with the high-performance data processing package data.table although, as documented in the issue Rdatatable/data.table#2273, is not fully compatible with sf objects.
Before using these capabilities, it is worth recapping how to discover the basic properties of vector data objects.
Let’s start by using base R functions to learn about the world dataset from the spData package:
class(world) # it's an sf object and a (tidy) data frame
#> [1] "sf" "tbl_df" "tbl" "data.frame"
dim(world) # it is a two-dimensional object, with 177 rows and 11 columns
#> [1] 177 11
world contains ten non-geographic columns (and one geometry list column) with almost 200 rows representing the world’s countries.
The function st_drop_geometry() keeps only the attributes data of an sf object, in other words removing its geometry:
world_df = st_drop_geometry(world)
class(world_df)
#> [1] "tbl_df" "tbl" "data.frame"
ncol(world_df)
#> [1] 10Dropping the geometry column before working with attribute data can be useful; data manipulation processes can run faster when they work only on the attribute data and geometry columns are not always needed.
For most cases, however, it makes sense to keep the geometry column, explaining why the column is ‘sticky’ (it remains after most attribute operations unless specifically dropped).
Non-spatial data operations on sf objects only change an object’s geometry when appropriate (e.g., by dissolving borders between adjacent polygons following aggregation).
Becoming skilled at geographic attribute data manipulation means becoming skilled at manipulating data frames.
For many applications, the tidyverse package dplyr (Wickham et al. 2023) offers an effective approach for working with data frames.
Tidyverse compatibility is an advantage of sf over its predecessor sp, but there are some pitfalls to avoid (see the supplementary tidyverse-pitfalls vignette at geocompx.org for details).
3.2.1 Vector attribute subsetting
Base R subsetting methods include the operator [ and the function subset().
The key dplyr subsetting functions are filter() and slice() for subsetting rows, and select() for subsetting columns.
Both approaches preserve the spatial components of attribute data in sf objects, while using the operator $ or the dplyr function pull() to return a single attribute column as a vector will lose the geometry data, as we will see.
This section focuses on subsetting sf data frames; for further details on subsetting vectors and non-geographic data frames we recommend reading Section 2.7 of An Introduction to R (R Core Team 2021) and chapter 4 of Advanced R Programming (Wickham 2019), respectively.
The [ operator can subset both rows and columns.
Indices placed inside square brackets placed directly after a data frame object name specify the elements to keep.
The command object[i, j] means ‘return the rows represented by i and the columns represented by j’, where i and j typically contain integers or TRUEs and FALSEs (indices can also be character strings, indicating row or column names).
object[5, 1:3], for example, means ’return data containing the 5th row and columns 1 to 3: the result should be a data frame with only 1 row and 3 columns, and a fourth geometry column if it’s an sf object.
Leaving i or j empty returns all rows or columns, so world[1:5, ] returns the first five rows and all 11 columns.
The examples below demonstrate subsetting with base R.
Guess the number of rows and columns in the sf data frames returned by each command and check the results on your own computer (see the end of the chapter for more exercises):
world[1:6, ] # subset rows by position
world[, 1:3] # subset columns by position
world[1:6, 1:3] # subset rows and columns by position
world[, c("name_long", "pop")] # columns by name
world[, c(T, T, F, F, F, F, F, T, T, F, F)] # by logical indices
world[, 888] # an index representing a non-existent columnA demonstration of the utility of using logical vectors for subsetting is shown in the code chunk below.
This creates a new object, small_countries, containing nations whose surface area is smaller than 10,000 km2.
i_small = world$area_km2 < 10000
summary(i_small) # a logical vector
#> Mode FALSE TRUE
#> logical 170 7
small_countries = world[i_small, ]The intermediary i_small (short for index representing small countries) is a logical vector that can be used to subset the seven smallest countries in the world by surface area.
A more concise command, which omits the intermediary object, generates the same result:
small_countries = world[world$area_km2 < 10000, ]The base R function subset() provides another way to achieve the same result:
small_countries = subset(world, area_km2 < 10000)
Base R functions are mature, stable and widely used, making them a rock solid choice, especially in contexts where reproducibility and reliability are key.
dplyr functions enable ‘tidy’ workflows which some people (the authors of this book included) find intuitive and productive for interactive data analysis, especially when combined with code editors such as RStudio that enable auto-completion of column names.
Key functions for subsetting data frames (including sf data frames) with dplyr functions are demonstrated below.
select() selects columns by name or position.
For example, you could select only two columns, name_long and pop, with the following command:
Note: as with the equivalent command in base R (world[, c("name_long", "pop")]), the sticky geom column remains.
select() also allows selecting a range of columns with the help of the : operator:
# all columns between name_long and pop (inclusive)
world2 = select(world, name_long:pop)You can remove specific columns with the - operator:
# all columns except subregion and area_km2 (inclusive)
world3 = select(world, -subregion, -area_km2)Subset and rename columns at the same time with the new_name = old_name syntax:
world4 = select(world, name_long, population = pop)It is worth noting that the command above is more concise than base R equivalent, which requires two lines of code:
world5 = world[, c("name_long", "pop")] # subset columns by name
names(world5)[names(world5) == "pop"] = "population" # rename column manuallyselect() also works with ‘helper functions’ for more advanced subsetting operations, including contains(), starts_with() and num_range() (see the help page with ?select for details).
Most dplyr verbs return a data frame, but you can extract a single column as a vector with pull().
You can get the same result in base R with the list subsetting operators $ and [[, the three following commands return the same numeric vector:
pull(world, pop)
world$pop
world[["pop"]]slice() is the row-equivalent of select().
The following code chunk, for example, selects rows 1 to 6:
slice(world, 1:6)filter() is dplyr’s equivalent of base R’s subset() function.
It keeps only rows matching given criteria, e.g., only countries with an area below a certain threshold, or with a high average of life expectancy, as shown in the following examples:
world7 = filter(world, area_km2 < 10000) # countries with a small area
world7 = filter(world, lifeExp > 82) # with high life expectancyThe standard set of comparison operators can be used in the filter() function, as illustrated in Table 3.1.
| Symbol | Name |
|---|---|
== |
Equal to |
!= |
Not equal to |
>, <
|
Greater/Less than |
>=, <=
|
Greater/Less than or equal |
&, |, !
|
Logical operators: And, Or, Not |
3.2.2 Chaining commands with pipes
Key to workflows using dplyr functions is the ‘pipe’ operator %>% (or since R 4.1.0 the native pipe |>), which takes its name from the Unix pipe | (Grolemund and Wickham 2016).
Pipes enable expressive code: the output of a previous function becomes the first argument of the next function, enabling chaining.
This is illustrated below, in which only countries from Asia are filtered from the world dataset, next the object is subset by columns (name_long and continent) and the first five rows (result not shown).
The above chunk shows how the pipe operator allows commands to be written in a clear order: the above run from top to bottom (line-by-line) and left to right. An alternative to piped operations is nested function calls, which are harder to read:
Another alternative is to split the operations into multiple self-contained lines, which is recommended when developing new R packages, an approach which has the advantage of saving intermediate results with distinct names which can be later inspected for debugging purposes (an approach which has disadvantages of being verbose and cluttering the global environment when undertaking interactive analysis):
world9_filtered = filter(world, continent == "Asia")
world9_selected = select(world9_filtered, continent)
world9 = slice(world9_selected, 1:5)Each approach has advantages and disadvantages, the importance of which depend on your programming style and applications. For interactive data analysis, the focus of this chapter, we find piped operations fast and intuitive, especially when combined with RStudio/VSCode shortcuts for creating pipes and auto-completing variable names.
3.2.3 Vector attribute aggregation
Aggregation involves summarizing data with one or more ‘grouping variables’, typically from columns in the data frame to be aggregated (geographic aggregation is covered in the next chapter).
An example of attribute aggregation is calculating the number of people per continent based on country-level data (one row per country).
The world dataset contains the necessary ingredients: the columns pop and continent, the population and the grouping variable, respectively.
The aim is to find the sum() of country populations for each continent, resulting in a smaller data frame (aggregation is a form of data reduction and can be a useful early step when working with large datasets).
This can be done with the base R function aggregate() as follows:
world_agg1 = aggregate(pop ~ continent, FUN = sum, data = world,
na.rm = TRUE)
class(world_agg1)
#> [1] "data.frame"The result is a non-spatial data frame with six rows, one per continent, and two columns reporting the name and population of each continent (see Table 3.2 with results for the top three most populous continents).
aggregate() is a generic function which means that it behaves differently depending on its inputs.
sf provides the method aggregate.sf() which is activated automatically when x is an sf object and a by argument is provided:
world_agg2 = aggregate(world["pop"], by = list(world$continent), FUN = sum,
na.rm = TRUE)
class(world_agg2)
#> [1] "sf" "data.frame"
nrow(world_agg2)
#> [1] 8The resulting world_agg2 object is a spatial object containing eight features representing the continents of the world (and the open ocean).
group_by() |> summarize() is the dplyr equivalent of aggregate().
Grouping variables are defined in the group_by() function and to aggregation formula is defined in the summarize() function, as shown below:
The approach may seem more complex, but it has benefits: flexibility, readability, and control over the new column names. This flexibility is illustrated in the command below, which calculates not only the population but also the area and number of countries in each continent:
world_agg4 = world |>
group_by(continent) |>
summarize(Pop = sum(pop, na.rm = TRUE), Area = sum(area_km2), N = n())In the previous code chunk Pop, Area and N are column names in the result, and sum() and n() were the aggregating functions.
These aggregating functions return sf objects with rows representing continents and geometries containing the multiple polygons representing each land mass and associated islands (this works thanks to the geometric operation ‘union’, as explained in Section 5.2.7).
Let’s combine what we have learned so far about dplyr functions, by chaining multiple commands to summarize attribute data about countries worldwide by continent.
The following command calculates population density (with mutate()), arranges continents by the number of countries they contain (with arrange()), and keeps only the three most populous continents (with slice_max()), the result of which is presented in Table 3.2):
world_agg5 = world |>
st_drop_geometry() |> # drop the geometry for speed
select(pop, continent, area_km2) |> # subset the columns of interest
group_by(Continent = continent) |> # group by continent and summarize:
summarize(Pop = sum(pop, na.rm = TRUE), Area = sum(area_km2), N = n()) |>
mutate(Density = round(Pop / Area)) |> # calculate population density
slice_max(Pop, n = 3) |> # keep only the top 3
arrange(desc(N)) # arrange in order of n. countries| Continent | Pop | Area | N | Density |
|---|---|---|---|---|
| Africa | 1154946633 | 29946198 | 51 | 39 |
| Asia | 4311408059 | 31252459 | 47 | 138 |
| Europe | 669036256 | 23065219 | 39 | 29 |
?summarize and vignette(package = "dplyr") and Chapter 5 of R for Data Science.
3.2.4 Vector attribute joining
Combining data from different sources is a common task in data preparation.
Joins do this by combining tables based on a shared ‘key’ variable.
dplyr has multiple join functions including left_join() and inner_join() — see vignette("two-table") for a full list.
These function names follow conventions used in the database language SQL (Grolemund and Wickham 2016, chap. 13); using them to join non-spatial datasets to sf objects is the focus of this section.
dplyr join functions work the same on data frames and sf objects, the only important difference being the geometry list column.
The result of data joins can be either an sf or data.frame object.
The most common type of attribute join on spatial data takes an sf object as the first argument and adds columns to it from a data.frame specified as the second argument.
To demonstrate joins, we will combine data on coffee production with the world dataset.
The coffee data is in a data frame called coffee_data from the spData package (see ?coffee_data for details).
It has three columns:
name_long names major coffee-producing nations and coffee_production_2016 and coffee_production_2017 contain estimated values for coffee production in units of 60-kg bags in each year.
A ‘left join’, which preserves the first dataset, merges world with coffee_data.
world_coffee = left_join(world, coffee_data)
#> Joining with `by = join_by(name_long)`
class(world_coffee)
#> [1] "sf" "tbl_df" "tbl" "data.frame"Because the input datasets share a ‘key variable’ (name_long) the join worked without using the by argument (see ?left_join for details).
The result is an sf object identical to the original world object but with two new variables (with column indices 11 and 12) on coffee production.
This can be plotted as a map, as illustrated in Figure 3.1, generated with the plot() function below.
names(world_coffee)
#> [1] "iso_a2" "name_long" "continent"
#> [4] "region_un" "subregion" "type"
#> [7] "area_km2" "pop" "lifeExp"
#> [10] "gdpPercap" "geom" "coffee_production_2016"
#> [13] "coffee_production_2017"
plot(world_coffee["coffee_production_2017"])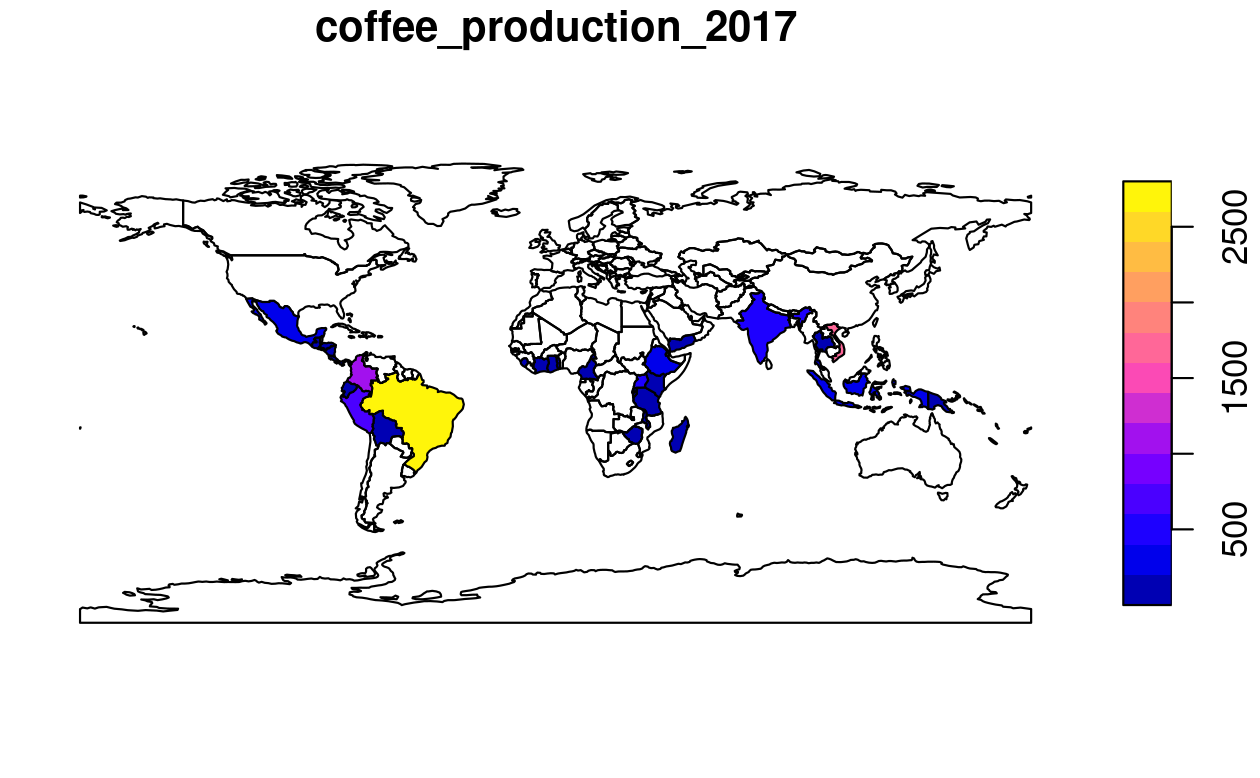
FIGURE 3.1: World coffee production (thousand 60-kg bags) by country, 2017. Source: International Coffee Organization.
For joining to work, a ‘key variable’ must be supplied in both datasets.
By default, dplyr uses all variables with matching names.
In this case, both coffee_data and world objects contained a variable called name_long, explaining the message Joining with 'by = join_by(name_long)'.
In the majority of cases where variable names are not the same, you have two options:
- Rename the key variable in one of the objects so they match.
- Use the
byargument to specify the joining variables.
The latter approach is demonstrated below on a renamed version of coffee_data.
coffee_renamed = rename(coffee_data, nm = name_long)
world_coffee2 = left_join(world, coffee_renamed, by = join_by(name_long == nm))Note that the name in the original object is kept, meaning that world_coffee and the new object world_coffee2 are identical.
Another feature of the result is that it has the same number of rows as the original dataset.
Although there are only 47 rows of data in coffee_data, all 177 country records are kept intact in world_coffee and world_coffee2:
rows in the original dataset with no match are assigned NA values for the new coffee production variables.
What if we only want to keep countries that have a match in the key variable?
In that case, an inner join can be used.
world_coffee_inner = inner_join(world, coffee_data)
#> Joining with `by = join_by(name_long)`
nrow(world_coffee_inner)
#> [1] 45Note that the result of inner_join() has only 45 rows compared with 47 in coffee_data.
What happened to the remaining rows?
We can identify the rows that did not match using the setdiff() function as follows:
setdiff(coffee_data$name_long, world$name_long)
#> [1] "Congo, Dem. Rep. of" "Others"The result shows that Others accounts for one row not present in the world dataset and that the name of the Democratic Republic of the Congo accounts for the other:
it has been abbreviated, causing the join to miss it.
The following command uses a string matching (regex) function from the stringr package to confirm what Congo, Dem. Rep. of should be.
drc = stringr::str_subset(world$name_long, "Dem*.+Congo")
drc
#> [1] "Democratic Republic of the Congo"To fix this issue, we will create a new version of coffee_data and update the name.
inner_join()ing the updated data frame returns a result with all 46 coffee-producing nations.
coffee_data$name_long[grepl("Congo,", coffee_data$name_long)] = drc
world_coffee_match = inner_join(world, coffee_data)
#> Joining with `by = join_by(name_long)`
nrow(world_coffee_match)
#> [1] 46It is also possible to join in the other direction: starting with a non-spatial dataset and adding variables from a simple features object.
This is demonstrated below, which starts with the coffee_data object and adds variables from the original world dataset.
In contrast with the previous joins, the result is not another simple feature object, but a data frame in the form of a tidyverse tibble:
the output of a join tends to match its first argument.
coffee_world = left_join(coffee_data, world)
#> Joining with `by = join_by(name_long)`
class(coffee_world)
#> [1] "tbl_df" "tbl" "data.frame"sf object.
The geometry column can only be used for creating maps and spatial operations if R ‘knows’ it is a spatial object, defined by a spatial package such as sf.
Fortunately, non-spatial data frames with a geometry list column (like coffee_world) can be coerced into an sf object as follows: st_as_sf(coffee_world).
This section covers the majority of joining use cases. For more information, we recommend reading the chapter Relational data in Grolemund and Wickham (2016), the join vignette in the geocompkg package that accompanies this book, and documentation describing joins with data.table and other packages. Additionally, spatial joins are covered in the next chapter (Section 4.2.5).
3.2.5 Creating attributes and removing spatial information
Often, we would like to create a new column based on already existing columns.
For example, we want to calculate population density for each country.
For this we need to divide a population column, here pop, by an area column, here area_km2 with unit area in square kilometers.
Using base R, we can type:
world_new = world # do not overwrite our original data
world_new$pop_dens = world_new$pop / world_new$area_km2
Alternatively, we can use one of dplyr functions: mutate() or transmute().
mutate() adds new columns at the penultimate position in the sf object (the last one is reserved for the geometry):
world_new2 = world |>
mutate(pop_dens = pop / area_km2)The difference between mutate() and transmute() is that the latter drops all other existing columns (except for the sticky geometry column).
unite() from the tidyr package (which provides many useful functions for reshaping datasets, including pivot_longer()) pastes together existing columns.
For example, we want to combine the continent and region_un columns into a new column named con_reg.
Additionally, we can define a separator (here, a colon :) which defines how the values of the input columns should be joined, and if the original columns should be removed (here, TRUE).
world_unite = world |>
tidyr::unite("con_reg", continent:region_un, sep = ":", remove = TRUE)The resulting sf object has a new column called con_reg representing the continent and region of each country, e.g., South America:Americas for Argentina and other South America countries.
tidyr’s separate() function does the opposite of unite(): it splits one column into multiple columns using either a regular expression or character positions.
The dplyr function rename() and the base R function setNames() are useful for renaming columns.
The first replaces an old name with a new one.
The following command, for example, renames the lengthy name_long column to simply name:
world |>
rename(name = name_long)
setNames() changes all column names at once, and requires a character vector with a name matching each column.
This is illustrated below, which outputs the same world object, but with very short names:
new_names = c("i", "n", "c", "r", "s", "t", "a", "p", "l", "gP", "geom")
world_new_names = world |>
setNames(new_names)
Each of these attribute data operations preserves the geometry of the simple features.
Sometimes, it makes sense to remove the geometry, for example to speed up aggregation.
Do this with st_drop_geometry(), not manually with commands such as select(world, -geom), as shown below.20
world_data = world |> st_drop_geometry()
class(world_data)
#> [1] "tbl_df" "tbl" "data.frame"3.3 Manipulating raster objects
In contrast to the vector data model underlying simple features (which represents points, lines and polygons as discrete entities in space), raster data represent continuous surfaces. This section shows how raster objects work by creating them from scratch, building on Section 2.3.2. Because of their unique structure, subsetting and other operations on raster datasets work in a different way, as demonstrated in Section 3.3.1.
The following code recreates the raster dataset used in Section 2.3.4, the result of which is illustrated in Figure 3.2.
This demonstrates how the rast() function works to create an example raster named elev (representing elevations).
elev = rast(nrows = 6, ncols = 6,
xmin = -1.5, xmax = 1.5, ymin = -1.5, ymax = 1.5,
vals = 1:36)The result is a raster object with 6 rows and 6 columns (specified by the nrow and ncol arguments), and a minimum and maximum spatial extent in x and y direction (xmin, xmax, ymin, ymax).
The vals argument sets the values that each cell contains: numeric data ranging from 1 to 36 in this case.
Raster objects can also contain categorical values of class logical or factor variables in R.
The following code creates the raster datasets shown in Figure 3.2:
grain_order = c("clay", "silt", "sand")
grain_char = sample(grain_order, 36, replace = TRUE)
grain_fact = factor(grain_char, levels = grain_order)
grain = rast(nrows = 6, ncols = 6,
xmin = -1.5, xmax = 1.5, ymin = -1.5, ymax = 1.5,
vals = grain_fact)
The raster object stores the corresponding look-up table or “Raster Attribute Table” (RAT) as a list of data frames, which can be viewed with cats(grain) (see ?cats() for more information).
Each element of this list is a layer of the raster.
It is also possible to use the function levels() for retrieving and adding new or replacing existing factor levels.
grain2 = grain # do not overwrite the original data
levels(grain2) = data.frame(value = c(0, 1, 2), wetness = c("wet", "moist", "dry"))
levels(grain2)
#> [[1]]
#> value wetness
#> 1 0 wet
#> 2 1 moist
#> 3 2 dry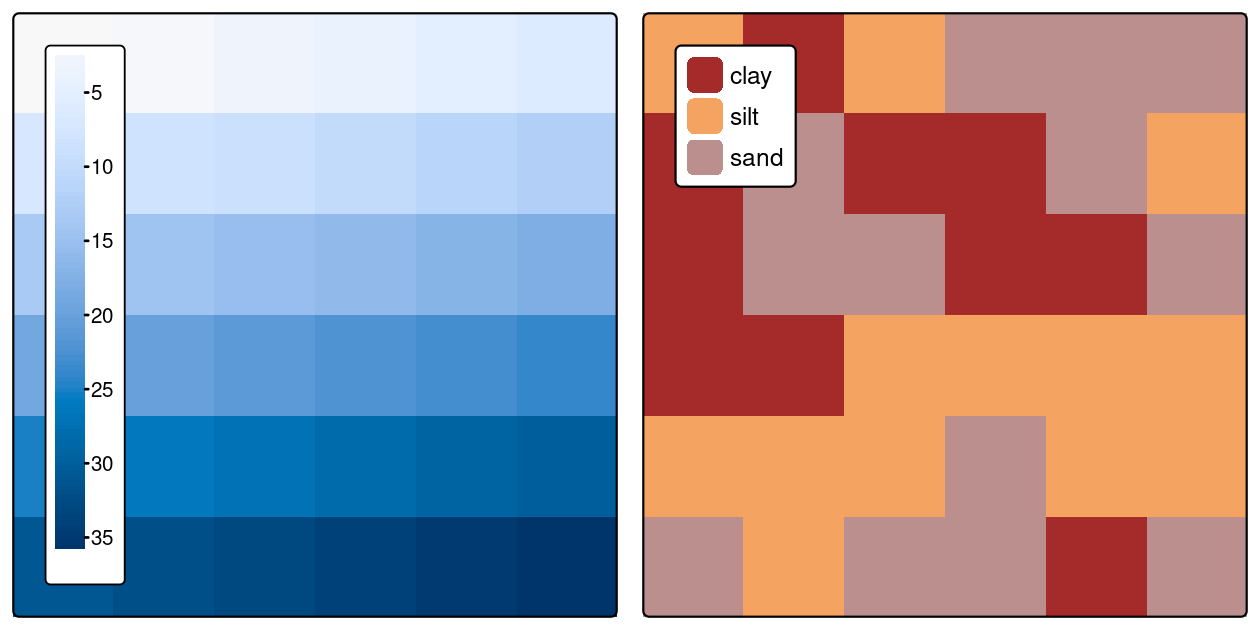
FIGURE 3.2: Raster datasets with numeric (left) and categorical values (right).
coltab() function (see ?coltab).
Importantly, saving a raster object with a color table to a file (e.g., GeoTIFF) will also save the color information.
3.3.1 Raster subsetting
Raster subsetting is done with the base R operator [, which accepts a variety of inputs:
- Row-column indexing
- Cell IDs
- Coordinates
- Another spatial object
Here, we only show the first two options since these can be considered non-spatial operations. If we need a spatial object to subset another or the output is a spatial object, we refer to this as spatial subsetting. Therefore, the latter two options will be shown in the next chapter (see Section 4.3.1).
The first two subsetting options are demonstrated in the commands below —
both return the value of the top left pixel in the raster object elev (results not shown).
# row 1, column 1
elev[1, 1]
# cell ID 1
elev[1]Subsetting of multi-layered raster objects will return the cell value(s) for each layer.
For example, two_layers = c(grain, elev); two_layers[1] returns a data frame with one row and two columns — one for each layer.
To extract all values, you can also use values().
Cell values can be modified by overwriting existing values in conjunction with a subsetting operation.
The following code chunk, for example, sets the upper left cell of elev to 0 (results not shown):
elev[1, 1] = 0
elev[]Leaving the square brackets empty is a shortcut version of values() for retrieving all values of a raster.
Multiple cells can also be modified in this way:
elev[1, c(1, 2)] = 0Replacing values of multi-layered rasters can be done with a matrix with as many columns as layers and rows as replaceable cells (results not shown):
3.3.2 Summarizing raster objects
terra contains functions for extracting descriptive statistics for entire rasters.
Printing a raster object to the console by typing its name returns minimum and maximum values of a raster.
summary() provides common descriptive statistics – minimum, maximum, quartiles and number of NAs for continuous rasters and the number of cells of each class for categorical rasters.
Further summary operations such as the standard deviation (see below) or custom summary statistics can be calculated with global().
global(elev, sd)summary() and global() functions with a multi-layered raster object, they will summarize each layer separately, as can be illustrated by running: summary(c(elev, grain)).
Additionally, the freq() function allows to get the frequency table of categorical values.
freq(grain)
#> layer value count
#> 1 1 clay 10
#> 2 1 silt 13
#> 3 1 sand 13Raster value statistics can be visualized in a variety of ways.
Specific functions such as boxplot(), density(), hist() and pairs() work also with raster objects, as demonstrated in the histogram created with the command below (not shown).
hist(elev)
In case the desired visualization function does not work with raster objects, one can extract the raster data to be plotted with the help of values() (Section 3.3.1).
Descriptive raster statistics belong to the so-called global raster operations. These and other typical raster processing operations are part of the map algebra scheme, which are covered in the next chapter (Section 4.3.2).
Some function names clash between packages (e.g., functions with the
name extract() exist in both terra and
tidyr packages). This may lead to unexpected results
when loading packages in a different order. In addition to calling
functions verbosely with their full namespace (e.g.,
tidyr::extract()) to avoid attaching packages with
library(), another way to prevent function name clashes is
by unloading the offending package with detach(). The
following command, for example, unloads the terra
package (this can also be done in the package tab which resides
by default in the right-bottom pane in RStudio):
detach(“package:terra”, unload = TRUE, force = TRUE). The
force argument makes sure that the package will be detached
even if other packages depend on it. This, however, may lead to a
restricted usability of packages depending on the detached package, and
it is therefore not recommended.
3.4 Exercises
For these exercises we will use the us_states and us_states_df datasets from the spData package.
You must have attached the package, and other packages used in the attribute operations chapter (sf, dplyr, terra) with commands such as library(spData) before attempting these exercises:
us_states is a spatial object (of class sf), containing geometry and a few attributes (including name, region, area, and population) of states within the contiguous United States.
us_states_df is a data frame (of class data.frame) containing the name and additional variables (including median income and poverty level, for the years 2010 and 2015) of US states, including Alaska, Hawaii and Puerto Rico.
The data comes from the United States Census Bureau, and is documented in ?us_states and ?us_states_df.
E1. Create a new object called us_states_name that contains only the NAME column from the us_states object using either base R ([) or tidyverse (select()) syntax.
What is the class of the new object and what makes it geographic?
E2. Select columns from the us_states object which contain population data.
Obtain the same result using a different command (bonus: try to find three ways of obtaining the same result).
Hint: try to use helper functions, such as contains or matches from dplyr (see ?contains).
E3. Find all states with the following characteristics (bonus: find and plot them):
- Belong to the Midwest region.
- Belong to the West region, have an area below 250,000 km2and in 2015 a population greater than 5,000,000 residents (hint: you may need to use the function
units::set_units()oras.numeric()). - Belong to the South region, had an area larger than 150,000 km2 and a total population in 2015 larger than 7,000,000 residents.
E4. What was the total population in 2015 in the us_states dataset?
What was the minimum and maximum total population in 2015?
E5. How many states are there in each region?
E6. What was the minimum and maximum total population in 2015 in each region? What was the total population in 2015 in each region?
E7. Add variables from us_states_df to us_states, and create a new object called us_states_stats.
What function did you use and why?
Which variable is the key in both datasets?
What is the class of the new object?
E8. us_states_df has two more rows than us_states.
How can you find them? (Hint: try to use the dplyr::anti_join() function.)
E9. What was the population density in 2015 in each state? What was the population density in 2010 in each state?
E10. How much has population density changed between 2010 and 2015 in each state? Calculate the change in percentages and map them.
E11. Change the columns’ names in us_states to lowercase. (Hint: helper functions - tolower() and colnames() may help.)
E12. Using us_states and us_states_df create a new object called us_states_sel.
The new object should have only two variables: median_income_15 and geometry.
Change the name of the median_income_15 column to Income.
E13. Calculate the change in the number of residents living below the poverty level between 2010 and 2015 for each state. (Hint: See ?us_states_df for documentation on the poverty level columns.) Bonus: Calculate the change in the percentage of residents living below the poverty level in each state.
E14. What was the minimum, average and maximum state’s number of people living below the poverty line in 2015 for each region? Bonus: What is the region with the largest increase in people living below the poverty line?
E15. Create a raster from scratch, with nine rows and columns and a resolution of 0.5 decimal degrees (WGS84). Fill it with random numbers. Extract the values of the four corner cells.
E16. What is the most common class of our example raster grain?
E17. Plot the histogram and the boxplot of the dem.tif file from the spDataLarge package (system.file("raster/dem.tif", package = "spDataLarge")).